- Paper: TIN-X: target importance and novelty explorer, Daniel C Cannon et al., Bioinformatics, 2017.
- Poster: TIN-X v2: modernized architecture with REST API for sustainability & interoperability, 2019.
The Target Importance and Novelty Explorer (TIN-X) is a web-based research tool for disease target prioritization. TIN-X prioritizes targets based on Novelty and Importance in the context of a chosen disease. Novelty is determined by the amount of publications related to a specific protein target with less publications correlating to greater novelty. Importance is determined by the amount of publications that study both the protein target and the chosen disease. Thus, TIN-X is a powerful tool providing researchers with the capability of quickly identifying protein targets that are correlated to a disease of interest and are poorly understood.
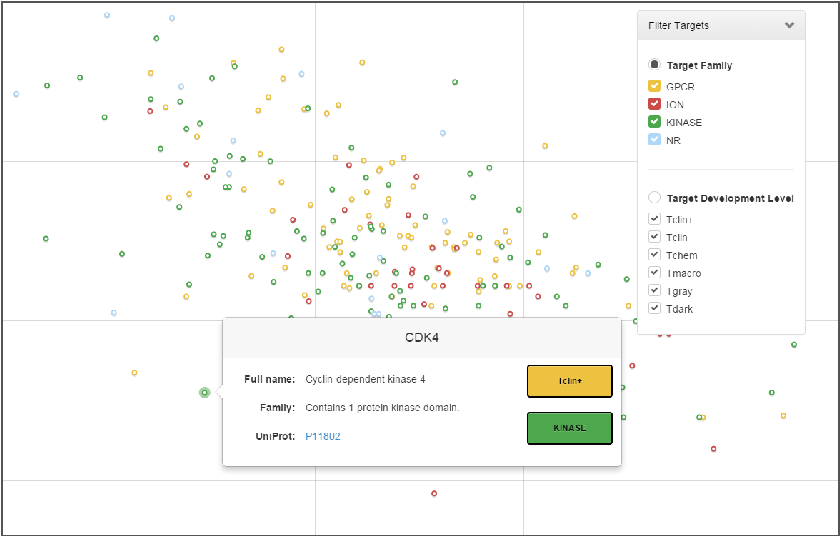
The TIN-X app provides a simple and intuitive interface to researchers who only have to choose the disease they are interested in studying from the list on the left-hand side of the screen (see Figure below). A scatter plot is then generated depicting Novelty along the X-Axis and Importance along the Y-Axis. Each point in the plot is a protein target associated with the chosen disease. These targets are color coded for quick target class identification and specific target details can be obtained by simply scrolling over the targets.
Researchers might be interested in targets in the different regions of the plot for various reasons. However, those targets along the upper right-hand side of the plot are probably the most interesting from the pharmaceutical perspective because these targets are poorly understood, but known to be relevant to the disease of interest.
Thus, TIN-X is an important research tool that will not only assist in providing researchers with greater accuracy in prioritizing targets, but also will save researchers time by helping them quickly identify interesting targets for further study and consequentially helping to illuminate poorly understood proteins worthy of more research.